OLS in R
Contents
OLS in R#
This document shows how to implement the topics we cover in cross-section econometrics in R.
# allows us to open stata datasets
library(foreign)
# for heteroskedasticity corrected variance/covariance matrices
library(sandwich)
# for White Heteroskedasticity Test
library(lmtest)
# for very pretty tables
library(stargazer)
# for bootstrapping
library(boot)
Load data and summarize:
tk.df = read.dta("http://rlhick.people.wm.edu/econ407/data/tobias_koop.dta")
tk4.df = subset(tk.df, time == 4)
stargazer(tk4.df,header=FALSE,title="Tobias and Koop Summary Statistics", type='text')
Tobias and Koop Summary Statistics
=================================================
Statistic N Mean St. Dev. Min Max
-------------------------------------------------
id 1,034 1,090.952 634.892 4 2,177
educ 1,034 12.275 1.567 9 19
ln_wage 1,034 2.138 0.466 0.420 3.590
pexp 1,034 4.815 2.190 0 12
time 1,034 4.000 0.000 4 4
ability 1,034 0.017 0.921 -3.140 1.890
meduc 1,034 11.403 3.027 0 20
feduc 1,034 11.585 3.736 0 20
broken_home 1,034 0.169 0.375 0 1
siblings 1,034 3.200 2.127 0 15
pexp2 1,034 27.980 22.599 0 144
-------------------------------------------------
Focusing on our dependent variable:
hist(tk4.df$ln_wage,main = "Log Wage")
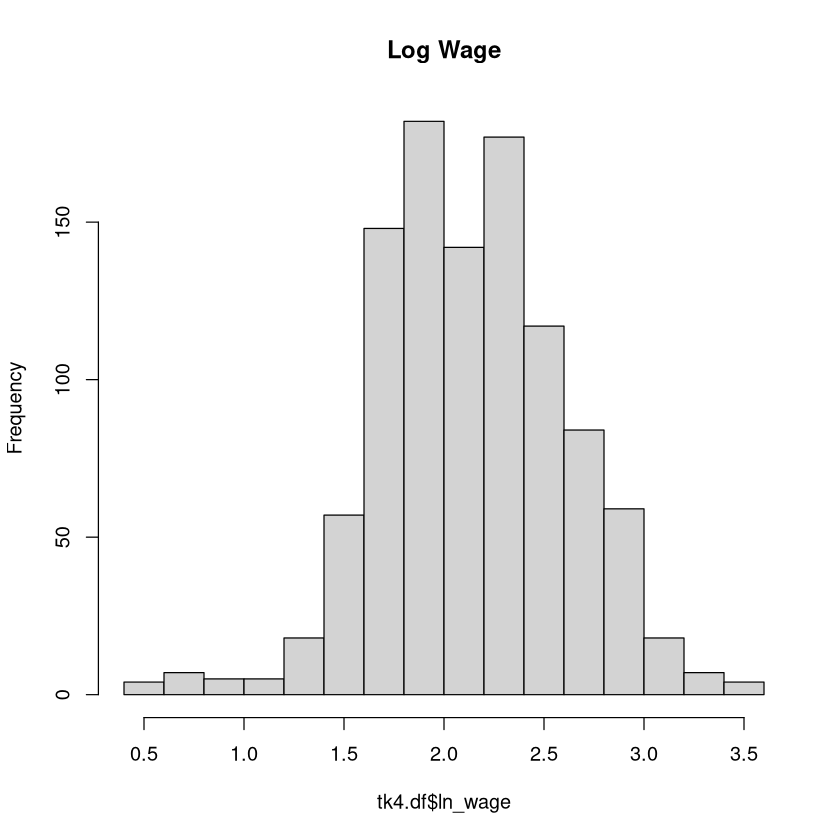
The OLS Model#
In R we use
ols.lm = lm(ln_wage~educ + pexp + pexp2 + broken_home, data=tk4.df)
summary(ols.lm)
Call:
lm(formula = ln_wage ~ educ + pexp + pexp2 + broken_home, data = tk4.df)
Residuals:
Min 1Q Median 3Q Max
-1.69490 -0.24818 -0.00079 0.25417 1.51139
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 0.460333 0.137294 3.353 0.000829 ***
educ 0.085273 0.009290 9.179 < 2e-16 ***
pexp 0.203521 0.023586 8.629 < 2e-16 ***
pexp2 -0.012413 0.002283 -5.438 6.73e-08 ***
broken_home -0.008725 0.035711 -0.244 0.807020
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
Residual standard error: 0.4265 on 1029 degrees of freedom
Multiple R-squared: 0.1664, Adjusted R-squared: 0.1632
F-statistic: 51.36 on 4 and 1029 DF, p-value: < 2.2e-16
Robust Standard Errors#
Note that since the \(\mathbf{b}\)’s are unbiased estimators for \(\beta\), those continue to be the ones to report. However, our variance covariance matrix is $\( VARCOV_{HC1} = \mathbf{(x'x)^{-1}\hat{V}(x'x)^{-1}} \)$
Defining \(\mathbf{e=y-xb}\), $\( \hat{\mathbf{V}} = diag(\mathbf{ee}') \)$
Implementing this in R, we can use the sandwich package’s vcovHC function with the HC1 option (stata default):
VARCOV = vcovHC(ols.lm, 'HC1')
se = sqrt(diag(VARCOV))
print(se)
(Intercept) educ pexp pexp2 broken_home
0.131541623 0.009929445 0.022683530 0.002244678 0.031238081
Note that the F-statistic reported by OLS is inappropriate in the presence of heteroskedasticity. We need to use a different form of the F test. Stata does this automatically. In R, we need to use waldtest for the correct model F statistic:
waldtest(ols.lm, vcov = vcovHC(ols.lm,'HC1'))
| Res.Df | Df | F | Pr(>F) | |
|---|---|---|---|---|
| <dbl> | <dbl> | <dbl> | <dbl> | |
| 1 | 1029 | NA | NA | NA |
| 2 | 1033 | -4 | 64.81968 | 6.411661e-49 |
We can put this together to make a more useful summary output:
# copy summary object
robust.summ = summary(ols.lm)
# overwrite all coefficients with robust counterparts
robust.summ$coefficients = unclass(coeftest(ols.lm, vcov. = VARCOV))
# overwrite f stat with correct robust value
f_robust = waldtest(ols.lm, vcov = VARCOV)
robust.summ$fstatistic[1] = f_robust$F[2]
print(robust.summ)
Call:
lm(formula = ln_wage ~ educ + pexp + pexp2 + broken_home, data = tk4.df)
Residuals:
Min 1Q Median 3Q Max
-1.69490 -0.24818 -0.00079 0.25417 1.51139
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 0.460333 0.131542 3.500 0.000486 ***
educ 0.085273 0.009929 8.588 < 2e-16 ***
pexp 0.203521 0.022684 8.972 < 2e-16 ***
pexp2 -0.012413 0.002245 -5.530 4.06e-08 ***
broken_home -0.008725 0.031238 -0.279 0.780057
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
Residual standard error: 0.4265 on 1029 degrees of freedom
Multiple R-squared: 0.1664, Adjusted R-squared: 0.1632
F-statistic: 64.82 on 4 and 1029 DF, p-value: < 2.2e-16
The Bootstrapping Approach#
In what follows below, I should you how to use R for bootstrapping regression coefficients, which will fix any heteroskedasticity problems we might have.
bs_ols <- function(formula, data, indices) {
d <- data[indices,] # allows boot to select sample
fit <- lm(formula, data=d)
return(coef(fit))
}
results <- boot(data=tk4.df, statistic=bs_ols,
R=1000, formula=ln_wage~educ + pexp + pexp2 + broken_home)
boot.ci(results,conf=0.95,type='perc',index=1)
boot.ci(results,conf=0.95,type='perc',index=2)
boot.ci(results,conf=0.95,type='perc',index=3)
boot.ci(results,conf=0.95,type='perc',index=4)
boot.ci(results,conf=0.95,type='perc',index=5)
colnames(results$t) = c("Intercept","educ","pexp","pexp2","broken_home")
summary(results$t)
BOOTSTRAP CONFIDENCE INTERVAL CALCULATIONS
Based on 1000 bootstrap replicates
CALL :
boot.ci(boot.out = results, conf = 0.95, type = "perc", index = 1)
Intervals :
Level Percentile
95% ( 0.2106, 0.7247 )
Calculations and Intervals on Original Scale
BOOTSTRAP CONFIDENCE INTERVAL CALCULATIONS
Based on 1000 bootstrap replicates
CALL :
boot.ci(boot.out = results, conf = 0.95, type = "perc", index = 2)
Intervals :
Level Percentile
95% ( 0.0645, 0.1035 )
Calculations and Intervals on Original Scale
BOOTSTRAP CONFIDENCE INTERVAL CALCULATIONS
Based on 1000 bootstrap replicates
CALL :
boot.ci(boot.out = results, conf = 0.95, type = "perc", index = 3)
Intervals :
Level Percentile
95% ( 0.1593, 0.2475 )
Calculations and Intervals on Original Scale
BOOTSTRAP CONFIDENCE INTERVAL CALCULATIONS
Based on 1000 bootstrap replicates
CALL :
boot.ci(boot.out = results, conf = 0.95, type = "perc", index = 4)
Intervals :
Level Percentile
95% (-0.0168, -0.0080 )
Calculations and Intervals on Original Scale
BOOTSTRAP CONFIDENCE INTERVAL CALCULATIONS
Based on 1000 bootstrap replicates
CALL :
boot.ci(boot.out = results, conf = 0.95, type = "perc", index = 5)
Intervals :
Level Percentile
95% (-0.0695, 0.0489 )
Calculations and Intervals on Original Scale
Intercept educ pexp pexp2
Min. :0.04771 Min. :0.05564 Min. :0.1316 Min. :-0.01988
1st Qu.:0.37305 1st Qu.:0.07856 1st Qu.:0.1877 1st Qu.:-0.01401
Median :0.46360 Median :0.08527 Median :0.2035 Median :-0.01236
Mean :0.46230 Mean :0.08513 Mean :0.2035 Mean :-0.01241
3rd Qu.:0.55261 3rd Qu.:0.09202 3rd Qu.:0.2196 3rd Qu.:-0.01081
Max. :0.84488 Max. :0.12228 Max. :0.2846 Max. :-0.00548
broken_home
Min. :-0.107363
1st Qu.:-0.029781
Median :-0.009216
Mean :-0.009060
3rd Qu.: 0.011689
Max. : 0.083809
Testing for Heteroskedasticity#
The two dominant heteroskedasticity tests are the White test and the Cook-Weisberg (both Breusch Pagan tests). The White test is particularly useful if you believe the error variances have a potentially non-linear relationship with model covariates.
In R we perform the White test by:
bptest(ols.lm,ln_wage ~ educ*pexp + educ*pexp2 + educ*broken_home +
pexp*pexp2 + pexp*broken_home + pexp2*broken_home +
I(educ^2) + I(pexp^2) + I(pexp2^2) + I(broken_home^2),
data=tk4.df)
studentized Breusch-Pagan test
data: ols.lm
BP = 29.196, df = 12, p-value = 0.003685
Notice that in R, we need to specify the form of the heteroskedasticity. To mirror the White test, we include interaction and second order terms. Both stata’s imtest and R’s bptest allow you to specify more esoteric forms of heteroskedasticity, although in R it is more apparent since we have to specify the form of the heteroskedasticity for the test to run.
A more basic test- one assuming that heteroskedastity is a linear function of the \(\mathbf{x}\)’s- is the Cook-Weisberg Breusch Pagan test. R doesn’t implement this test directly. But it can be performed manually following the steps we discussed in class:
Recover residuals as above, denote as \(\mathbf{e}\)
square the residuals by the sqare root of mean squared residuals: $\( \mathbf{r} = \frac{diagonal(\mathbf{ee}')}{\sqrt{\mathbf{e'e}/N}} \)$
Run the regression \(\mathbf{r} = \gamma_0 + \gamma_1 \mathbf{xb} + \mathbf{v}\). Recall that \(\mathbf{xb} = \mathbf{\hat{y}}\).
Recover ESS defined as $\( ESS = (\mathbf{\hat{r}} - \bar{r})'(\hat{\mathbf{r}} - \bar{r}) \)$
ESS/2 is the test statistic distributed \(\chi^2_{(1)}\)